Node = "Manager", parm = list("rotat",NA)) # The first element in parm can be a regular expression # It is possible to select unique managements, but not non-unique ones # This has many options and a complex structure Inspect_apsim("maize-soybean-rotation.apsim", src.dir = extd.dir, node = "Crop") Inspect_apsim("maize-soybean-rotation.apsim", src.dir = extd.dir, Inspect_apsim("maize-soybean-rotation.apsim", src.dir = extd.dir, node = "Soil", Inspect_apsim("maize-soybean-rotation.apsim", src.dir = extd.dir, node = "Weather") Inspect_apsim("maize-soybean-rotation.apsim", src.dir = extd.dir, node = "Clock") # Testing with maize-soybean-rotation.apsim Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Outputfile", Soil.child = "Water", parm = list("Barley", "LL"), # 'parm' can be a list or a character vector of length equal to two # but still there will be multiple soil layers # To print the parm.path the selection needs to be unique Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Soil", # when soil.child = "Water" there are potentially many crops to chose from Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Crop", parm = list("sow",7)) Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Crop", parm = list("sow",NA)) Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "SurfaceOrganicMatter") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Soil", soil.child = "Sample") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Soil", soil.child = "InitialWater") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Soil", soil.child = "Analysis") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Soil", soil.child = "OrganicMatter") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Soil", soil.child = "Metadata") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Weather") Inspect_apsim("Millet.apsim", src.dir = extd.dir, node = "Clock") In this case the printed path willĮxtd.dir <- system.file("extdata", package = "apsimx")
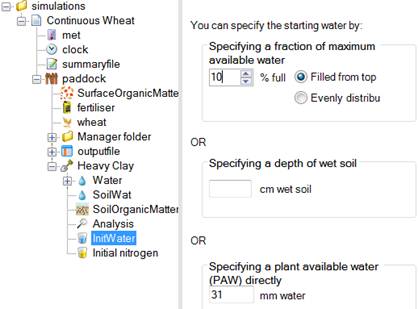
(or a regular expression), the appropriate node can be selected. By providing the name of the folder to root

The instance in the first folder unless 'root' is supplied. When multiple folders are present as it is the case when there are factorials. It returns the parameter path (when print.path equals TRUE) and table with inspected parameters and values
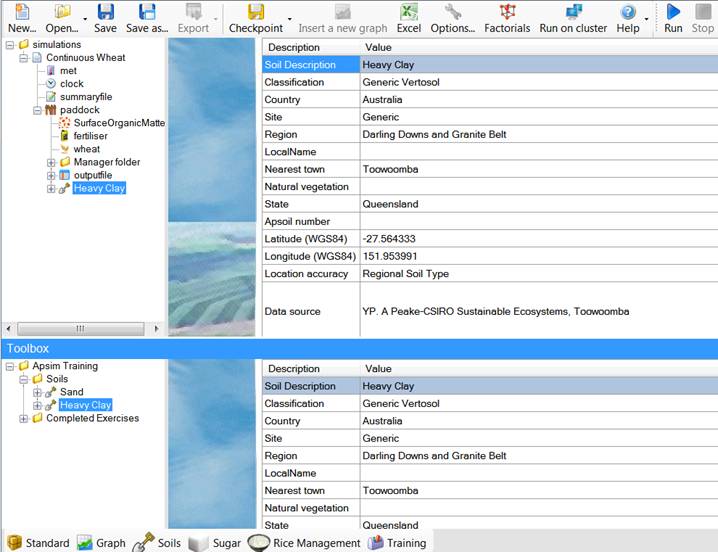
apsim file.įor ‘Crop’, ‘Manager’ and ‘Other’, ‘parm’ should be indicated with a first element to look for and a second with the relative position in case there are It does not print all information from an. This is simply a script that prints the relevant parameters which are likely to need editing. This can be specified by supplying an appropriate character. In simulation structures such as factorials there will be multiple possible nodes. Whether to print the parameter path (default = FALSE) Parameter to inspect when node = ‘Crop’, ‘Manager’, ‘Outputfile’ or ‘Other’ ‘MicroClimate’, ‘Crop’, ‘Manager’, ‘Outputfile’ or ‘Other’ apsim file to be inspected defaults to the current working directoryĮither ‘Weather’, ‘Soil’, ‘SurfaceOrganicMatter’, apsim (Classic) to be inspected (XML)ĭirectory containing the. Soil.child = c("Metadata", "Water", "OrganicMatter", "Nitrogen", "Analysis",įile ending in. Node = c("Clock", "Weather", "Soil", "SurfaceOrganicMatter", "Crop", "Manager", It does not replace the GUI, but it can save time by quickly checking parameters and values.
